2D data regression: Python Ordinary Kriging interpolation code
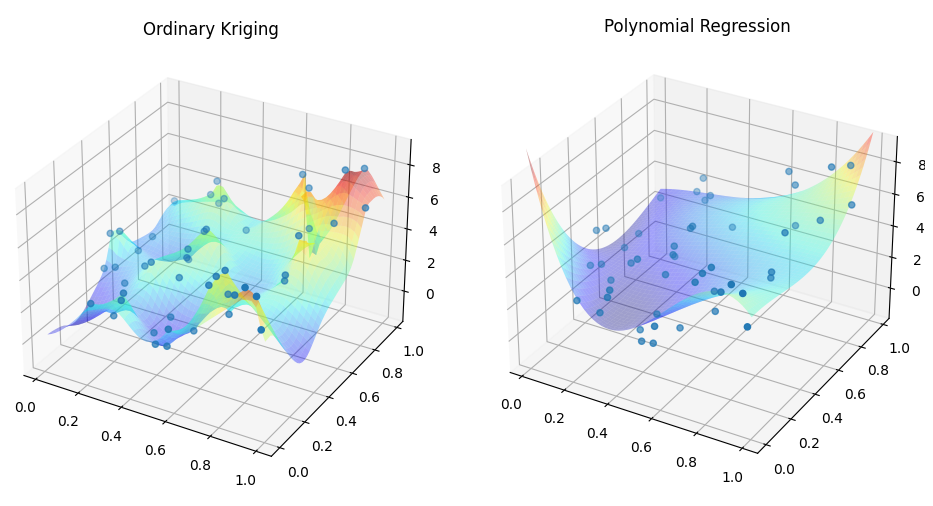
Ordinary Kriging is one of the technique to build data-exact, best linear unbiased estimator. Goal here is to write a code to perform ordinary kriging using 2-D simple dataset, then compare the results with simple 3rd order polynomial function.
First, results obtained below code are shown below.
import itertools
import os
import numpy as np
import matplotlib.cm as cm
import matplotlib.mlab as mlab
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
def main():
pass
def Test_Okriging(x,y,Obsp,xx,yy,nget,sill,rnge):
nobs = len(Obsp)
A = np.ones((nobs+1,nobs+1)) # = 1 for lagrange multiplier
b = np.ones((nobs+1,1))
Cd = np.ones((nobs+1,nobs+1))
b[nobs]= 1 # 1 = lagrange multiple
pos = np.ones((nobs,2))
pos[:,0] = x/np.max(x)
pos[:,1] = y/np.max(y)
# Variogram_Generation(Cd,A,Obsp,pos,nget,sill,rnge)
#------------------------------------------------
# Covariance Generation by Data-Data Distance
for i in range(nobs):
for j in range(i, nobs) :
Cd[i][j] = np.linalg.norm(pos[i]-pos[j])
#------------------------------------------------
#------------------------------------------------
# Variogram: Spherical Method
for i in range(nobs) :
for j in range(i, nobs) :
if Cd[i][j] < rnge :
A[i][j] = A[j][i] = nget + sill*(1.5*Cd[i][j]/rnge - 0.5*(Cd[i][j]/rnge)**3)
else:
A[i][j] = A[j][i] = nget + sill
#------------------------------------------------
#---------initialize values--------
posidim = len(pos[0])
mesh = len(xx)
cnt_x = (1. - 0.)/mesh
cnt_y = (1. - 0.)/mesh
vvval = np.ones((int(mesh),int(mesh)))
#--------- estimate all location --------
cord = np.ones((posidim))
for i in range(mesh):
cord[0] = cnt_x*i+0.
for j in range(mesh):
cord[1] = cnt_y*j+0.
cord[0] = xx[i,j]
cord[1] = yy[i,j]
#---Current to Obs Distance Yo ------------------
for k in range(nobs) :
distance = np.linalg.norm(cord-pos[k])
if distance < rnge :
b[k] = nget + sill*(1.5*distance/rnge - 0.5*(distance/rnge)**3)
else:
b[k] = nget + sill
#-------------------------------
Weit = linalg.solve(A,b)
OKest = np.sum([Weit[i]*Obsp[i] for i in range(0, nobs-1)])
vvval[i,j] = OKest
#--------- Return! ---------
return vvval
#-------------------------------------------------------------------
# Function for polynomial 2D
def polyfit2d(x, y, z, order):
ncols = (order + 1)**2
G = np.zeros((x.size, ncols))
ij = itertools.product(range(order+1), range(order+1))
for k, (i,j) in enumerate(ij):
G[:,k] = x**i * y**j
m, _, _, _ = np.linalg.lstsq(G, z)
return m
#-------------------------------------------------------------------
# Function for polynomial 2D
def polyval2d(x, y, m):
order = int(np.sqrt(len(m))) - 1
ij = itertools.product(range(order+1), range(order+1))
z = np.zeros_like(x)
for a, (i,j) in zip(m, ij):
z += a * x**i * y**j
return z
#-------------------------------------------------------------------
# Function main
#-------------------------------------------------------------------
if __name__ == '__main__':
main()
from numpy import matrix
from numpy import linalg
from scipy.interpolate import Rbf
#------------------------------------------
# Get polynomial Coefficient : 2D, Random
numdata = 50
np.random.seed(65537)
x = np.random.random(numdata)
y = np.random.random(numdata)
xy = np.ones((numdata,2))
xy[:,0] = x/np.max(x)
xy[:,1] = y/np.max(y)
z = x**2 + y**2 + 3*x**3 + y + np.random.random(numdata)*4 # Noise
#------------------------------------------
# Get polynomial Coefficient : 2D, Random
mesh = 50
xx, yy = np.meshgrid(np.linspace(x.min(), x.max(), mesh), np.linspace(y.min(), y.max(), mesh))
#------------------------------------------
# Ordinary Kriging Estimation
#------------------------------------------
zz = Test_Okriging(x,y,z,xx,yy,0.0,1.0,0.35)
fig = plt.figure()
ax = fig.add_subplot(121, projection='3d')
ax.scatter(x, y, z)
ax.plot_surface(xx, yy, zz,rstride = 1, cstride = 1, alpha=0.35, cmap=cm.jet)
plt.title("Ordinary Kriging");
ax.set_zlim3d(-z.min()-1, z.max()+1)
#------------------------------------------
# Polynomial fitting
#------------------------------------------
m = polyfit2d(x,y,z,3) # Fit 2d polynomial
zz = polyval2d(xx, yy, m) # Meshing by polynomial
#fig = plt.figure()
ax = fig.add_subplot(122, projection='3d')
ax.scatter(x, y, z)
ax.plot_surface(xx, yy, zz, rstride=1, cstride=1, alpha=0.35, cmap=cm.jet)
plt.title("Polynomial Regression");
ax.set_zlim3d(-z.min()-1, z.max()+1)
plt.show()
import itertools
import os
import numpy as np
import matplotlib.cm as cm
import matplotlib.mlab as mlab
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
def main():
pass
def Test_Okriging(x,y,Obsp,xx,yy,nget,sill,rnge):
nobs = len(Obsp)
A = np.ones((nobs+1,nobs+1)) # = 1 for lagrange multiplier
b = np.ones((nobs+1,1))
Cd = np.ones((nobs+1,nobs+1))
b[nobs]= 1 # 1 = lagrange multiple
pos = np.ones((nobs,2))
pos[:,0] = x/np.max(x)
pos[:,1] = y/np.max(y)
# Variogram_Generation(Cd,A,Obsp,pos,nget,sill,rnge)
#------------------------------------------------
# Covariance Generation by Data-Data Distance
for i in range(nobs):
for j in range(i, nobs) :
Cd[i][j] = np.linalg.norm(pos[i]-pos[j])
#------------------------------------------------
#------------------------------------------------
# Variogram: Spherical Method
for i in range(nobs) :
for j in range(i, nobs) :
if Cd[i][j] < rnge :
A[i][j] = A[j][i] = nget + sill*(1.5*Cd[i][j]/rnge - 0.5*(Cd[i][j]/rnge)**3)
else:
A[i][j] = A[j][i] = nget + sill
#------------------------------------------------
#---------initialize values--------
posidim = len(pos[0])
mesh = len(xx)
cnt_x = (1. - 0.)/mesh
cnt_y = (1. - 0.)/mesh
vvval = np.ones((int(mesh),int(mesh)))
#--------- estimate all location --------
cord = np.ones((posidim))
for i in range(mesh):
cord[0] = cnt_x*i+0.
for j in range(mesh):
cord[1] = cnt_y*j+0.
cord[0] = xx[i,j]
cord[1] = yy[i,j]
#---Current to Obs Distance Yo ------------------
for k in range(nobs) :
distance = np.linalg.norm(cord-pos[k])
if distance < rnge :
b[k] = nget + sill*(1.5*distance/rnge - 0.5*(distance/rnge)**3)
else:
b[k] = nget + sill
#-------------------------------
Weit = linalg.solve(A,b)
OKest = np.sum([Weit[i]*Obsp[i] for i in range(0, nobs-1)])
vvval[i,j] = OKest
#--------- Return! ---------
return vvval
#-------------------------------------------------------------------
# Function for polynomial 2D
def polyfit2d(x, y, z, order):
ncols = (order + 1)**2
G = np.zeros((x.size, ncols))
ij = itertools.product(range(order+1), range(order+1))
for k, (i,j) in enumerate(ij):
G[:,k] = x**i * y**j
m, _, _, _ = np.linalg.lstsq(G, z)
return m
#-------------------------------------------------------------------
# Function for polynomial 2D
def polyval2d(x, y, m):
order = int(np.sqrt(len(m))) - 1
ij = itertools.product(range(order+1), range(order+1))
z = np.zeros_like(x)
for a, (i,j) in zip(m, ij):
z += a * x**i * y**j
return z
#-------------------------------------------------------------------
# Function main
#-------------------------------------------------------------------
if __name__ == '__main__':
main()
from numpy import matrix
from numpy import linalg
from scipy.interpolate import Rbf
#------------------------------------------
# Get polynomial Coefficient : 2D, Random
numdata = 50
np.random.seed(65537)
x = np.random.random(numdata)
y = np.random.random(numdata)
xy = np.ones((numdata,2))
xy[:,0] = x/np.max(x)
xy[:,1] = y/np.max(y)
z = x**2 + y**2 + 3*x**3 + y + np.random.random(numdata)*4 # Noise
#------------------------------------------
# Get polynomial Coefficient : 2D, Random
mesh = 50
xx, yy = np.meshgrid(np.linspace(x.min(), x.max(), mesh), np.linspace(y.min(), y.max(), mesh))
#------------------------------------------
# Ordinary Kriging Estimation
#------------------------------------------
zz = Test_Okriging(x,y,z,xx,yy,0.0,1.0,0.35)
fig = plt.figure()
ax = fig.add_subplot(121, projection='3d')
ax.scatter(x, y, z)
ax.plot_surface(xx, yy, zz,rstride = 1, cstride = 1, alpha=0.35, cmap=cm.jet)
plt.title("Ordinary Kriging");
ax.set_zlim3d(-z.min()-1, z.max()+1)
#------------------------------------------
# Polynomial fitting
#------------------------------------------
m = polyfit2d(x,y,z,3) # Fit 2d polynomial
zz = polyval2d(xx, yy, m) # Meshing by polynomial
#fig = plt.figure()
ax = fig.add_subplot(122, projection='3d')
ax.scatter(x, y, z)
ax.plot_surface(xx, yy, zz, rstride=1, cstride=1, alpha=0.35, cmap=cm.jet)
plt.title("Polynomial Regression");
ax.set_zlim3d(-z.min()-1, z.max()+1)
plt.show()
import itertools
import os
import numpy as np
import matplotlib.cm as cm
import matplotlib.mlab as mlab
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
def main():
pass
def Test_Okriging(x,y,Obsp,xx,yy,nget,sill,rnge):
nobs = len(Obsp)
A = np.ones((nobs+1,nobs+1)) # = 1 for lagrange multiplier
b = np.ones((nobs+1,1))
Cd = np.ones((nobs+1,nobs+1))
b[nobs]= 1 # 1 = lagrange multiple
pos = np.ones((nobs,2))
pos[:,0] = x/np.max(x)
pos[:,1] = y/np.max(y)
# Variogram_Generation(Cd,A,Obsp,pos,nget,sill,rnge)
#------------------------------------------------
# Covariance Generation by Data-Data Distance
for i in range(nobs):
for j in range(i, nobs) :
Cd[i][j] = np.linalg.norm(pos[i]-pos[j])
#------------------------------------------------
#------------------------------------------------
# Variogram: Spherical Method
for i in range(nobs) :
for j in range(i, nobs) :
if Cd[i][j] < rnge :
A[i][j] = A[j][i] = nget + sill*(1.5*Cd[i][j]/rnge - 0.5*(Cd[i][j]/rnge)**3)
else:
A[i][j] = A[j][i] = nget + sill
#------------------------------------------------
#---------initialize values--------
posidim = len(pos[0])
mesh = len(xx)
cnt_x = (1. - 0.)/mesh
cnt_y = (1. - 0.)/mesh
vvval = np.ones((int(mesh),int(mesh)))
#--------- estimate all location --------
cord = np.ones((posidim))
for i in range(mesh):
cord[0] = cnt_x*i+0.
for j in range(mesh):
cord[1] = cnt_y*j+0.
cord[0] = xx[i,j]
cord[1] = yy[i,j]
#---Current to Obs Distance Yo ------------------
for k in range(nobs) :
distance = np.linalg.norm(cord-pos[k])
if distance < rnge :
b[k] = nget + sill*(1.5*distance/rnge - 0.5*(distance/rnge)**3)
else:
b[k] = nget + sill
#-------------------------------
Weit = linalg.solve(A,b)
OKest = np.sum([Weit[i]*Obsp[i] for i in range(0, nobs-1)])
vvval[i,j] = OKest
#--------- Return! ---------
return vvval
#-------------------------------------------------------------------
# Function for polynomial 2D
def polyfit2d(x, y, z, order):
ncols = (order + 1)**2
G = np.zeros((x.size, ncols))
ij = itertools.product(range(order+1), range(order+1))
for k, (i,j) in enumerate(ij):
G[:,k] = x**i * y**j
m, _, _, _ = np.linalg.lstsq(G, z)
return m
#-------------------------------------------------------------------
# Function for polynomial 2D
def polyval2d(x, y, m):
order = int(np.sqrt(len(m))) - 1
ij = itertools.product(range(order+1), range(order+1))
z = np.zeros_like(x)
for a, (i,j) in zip(m, ij):
z += a * x**i * y**j
return z
#-------------------------------------------------------------------
# Function main
#-------------------------------------------------------------------
if __name__ == '__main__':
main()
from numpy import matrix
from numpy import linalg
from scipy.interpolate import Rbf
#------------------------------------------
# Get polynomial Coefficient : 2D, Random
numdata = 50
np.random.seed(65537)
x = np.random.random(numdata)
y = np.random.random(numdata)
xy = np.ones((numdata,2))
xy[:,0] = x/np.max(x)
xy[:,1] = y/np.max(y)
z = x**2 + y**2 + 3*x**3 + y + np.random.random(numdata)*4 # Noise
#------------------------------------------
# Get polynomial Coefficient : 2D, Random
mesh = 50
xx, yy = np.meshgrid(np.linspace(x.min(), x.max(), mesh), np.linspace(y.min(), y.max(), mesh))
#------------------------------------------
# Ordinary Kriging Estimation
#------------------------------------------
zz = Test_Okriging(x,y,z,xx,yy,0.0,1.0,0.35)
fig = plt.figure()
ax = fig.add_subplot(121, projection='3d')
ax.scatter(x, y, z)
ax.plot_surface(xx, yy, zz,rstride = 1, cstride = 1, alpha=0.35, cmap=cm.jet)
plt.title("Ordinary Kriging");
ax.set_zlim3d(-z.min()-1, z.max()+1)
#------------------------------------------
# Polynomial fitting
#------------------------------------------
m = polyfit2d(x,y,z,3) # Fit 2d polynomial
zz = polyval2d(xx, yy, m) # Meshing by polynomial
#fig = plt.figure()
ax = fig.add_subplot(122, projection='3d')
ax.scatter(x, y, z)
ax.plot_surface(xx, yy, zz, rstride=1, cstride=1, alpha=0.35, cmap=cm.jet)
plt.title("Polynomial Regression");
ax.set_zlim3d(-z.min()-1, z.max()+1)
plt.show()